Calcium »
PDB 3fil-3fye »
3fye »
Calcium in PDB 3fye: Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State
Enzymatic activity of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State
All present enzymatic activity of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State:
1.9.3.1;
1.9.3.1;
Protein crystallography data
The structure of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State, PDB code: 3fye
was solved by
L.Qin,
D.A.Mills,
D.A.Proshlyakov,
C.Hiser,
S.Ferguson-Miller,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 50.00 / 2.15 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 124.613, 131.472, 176.192, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.6 / 22.1 |
Other elements in 3fye:
The structure of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State also contains other interesting chemical elements:
| Magnesium | (Mg) | 2 atoms |
| Cadmium | (Cd) | 4 atoms |
| Iron | (Fe) | 4 atoms |
| Copper | (Cu) | 6 atoms |
Calcium Binding Sites:
The binding sites of Calcium atom in the Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State
(pdb code 3fye). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total 2 binding sites of Calcium where determined in the Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State, PDB code: 3fye:
Jump to Calcium binding site number: 1; 2;
In total 2 binding sites of Calcium where determined in the Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State, PDB code: 3fye:
Jump to Calcium binding site number: 1; 2;
Calcium binding site 1 out of 2 in 3fye
Go back to
Calcium binding site 1 out
of 2 in the Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State
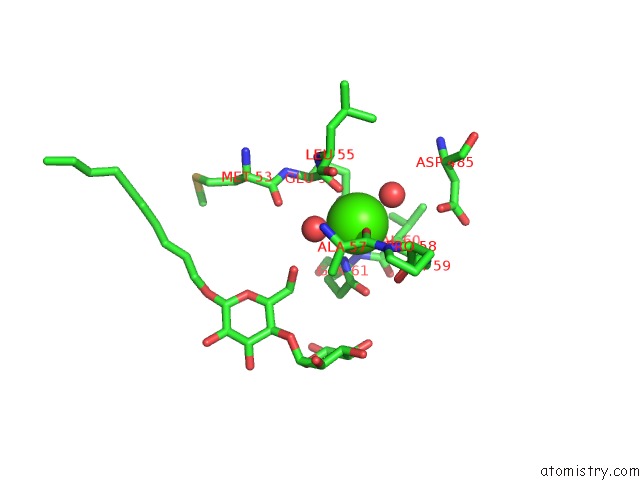
Mono view
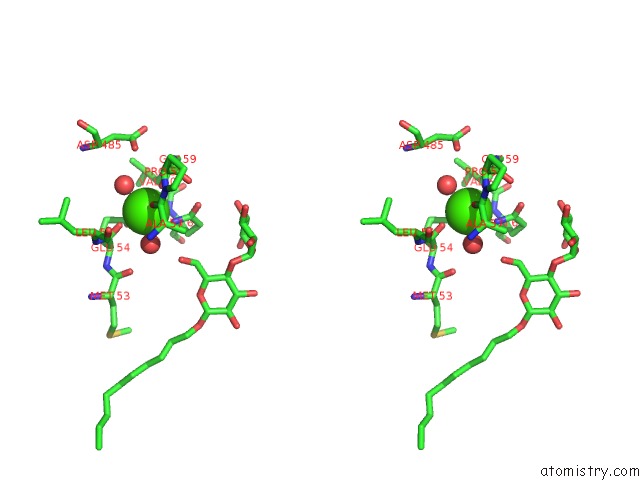
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State within 5.0Å range:
|
Calcium binding site 2 out of 2 in 3fye
Go back to
Calcium binding site 2 out
of 2 in the Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State
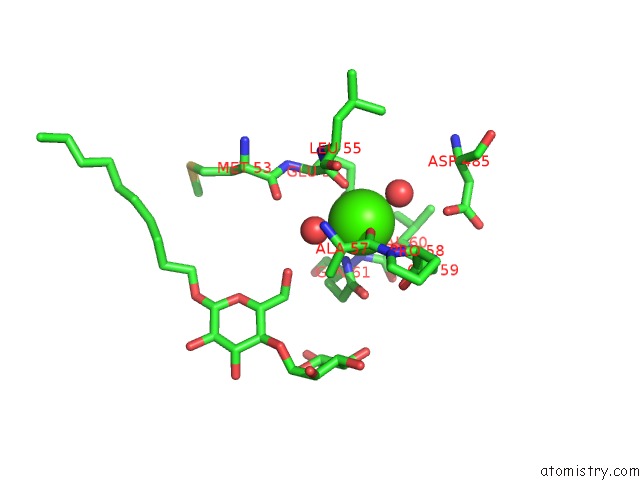
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 2 of Catalytic Core Subunits (I and II) of Cytochrome C Oxidase From Rhodobacter Sphaeroides in the Reduced State within 5.0Å range:
|
Reference:
L.Qin,
J.Liu,
D.Mills,
D.A.Proshlyakov,
C.Hiser,
S.Ferguson-Miller.
Redox Dependent Conformational Changes in Cytochrome C Oxidase Suggest A Gating Mechanism For Proton Uptake. Biochemistry V. 48 5121 2009.
ISSN: ISSN 0006-2960
PubMed: 19397279
DOI: 10.1021/BI9001387
Page generated: Sat Jul 13 10:21:51 2024
ISSN: ISSN 0006-2960
PubMed: 19397279
DOI: 10.1021/BI9001387
Last articles
Ca in 3FNLCa in 3FMG
Ca in 3FM6
Ca in 3FLT
Ca in 3FM4
Ca in 3FM1
Ca in 3FLR
Ca in 3FL8
Ca in 3FL9
Ca in 3FLF