Calcium »
PDB 5szm-5tad »
5t9c »
Calcium in PDB 5t9c: Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak)
Enzymatic activity of Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak)
All present enzymatic activity of Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak):
3.1.4.46;
3.1.4.46;
Protein crystallography data
The structure of Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak), PDB code: 5t9c
was solved by
F.K.K.Li,
N.C.J.Strynadka,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 48.49 / 1.48 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 47.101, 58.240, 87.565, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16.8 / 19.7 |
Other elements in 5t9c:
The structure of Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak) also contains other interesting chemical elements:
| Sodium | (Na) | 1 atom |
Calcium Binding Sites:
The binding sites of Calcium atom in the Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak)
(pdb code 5t9c). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total only one binding site of Calcium was determined in the Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak), PDB code: 5t9c:
In total only one binding site of Calcium was determined in the Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak), PDB code: 5t9c:
Calcium binding site 1 out of 1 in 5t9c
Go back to
Calcium binding site 1 out
of 1 in the Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak)
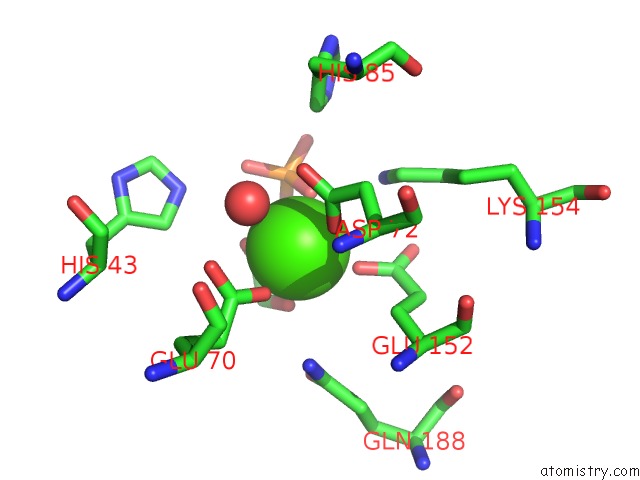
Mono view
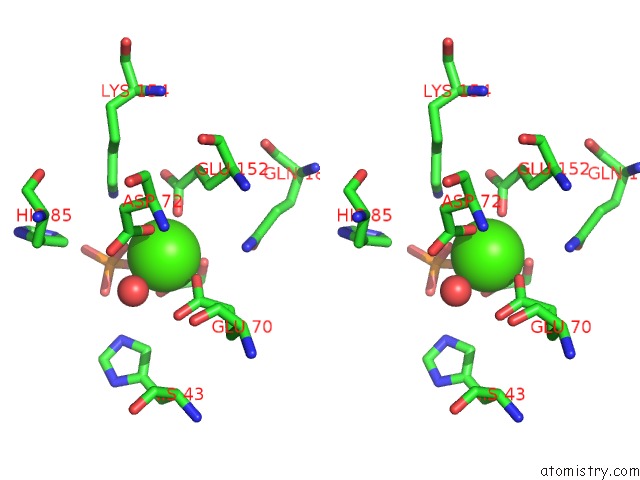
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Crystal Structure of B. Subtilis 168 Glpq in Complex with Glycerol-3- Phosphate (1 Hour Soak) within 5.0Å range:
|
Reference:
C.L.Myers,
F.K.Li,
B.M.Koo,
O.M.El-Halfawy,
S.French,
C.A.Gross,
N.C.Strynadka,
E.D.Brown.
Identification of Two Phosphate Starvation-Induced Wall Teichoic Acid Hydrolases Provides First Insights Into the Degradative Pathway of A Key Bacterial Cell Wall Component. J. Biol. Chem. V. 291 26066 2016.
ISSN: ESSN 1083-351X
PubMed: 27780866
DOI: 10.1074/JBC.M116.760447
Page generated: Wed Jul 9 10:08:56 2025
ISSN: ESSN 1083-351X
PubMed: 27780866
DOI: 10.1074/JBC.M116.760447
Last articles
Fe in 2YXOFe in 2YRS
Fe in 2YXC
Fe in 2YNM
Fe in 2YVJ
Fe in 2YP1
Fe in 2YU2
Fe in 2YU1
Fe in 2YQB
Fe in 2YOO