Calcium »
PDB 6d7p-6dlh »
6dbl »
Calcium in PDB 6dbl: Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
Enzymatic activity of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
All present enzymatic activity of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas:
2.3.2.27;
2.3.2.27;
Other elements in 6dbl:
The structure of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
Calcium Binding Sites:
The binding sites of Calcium atom in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
(pdb code 6dbl). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total 4 binding sites of Calcium where determined in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas, PDB code: 6dbl:
Jump to Calcium binding site number: 1; 2; 3; 4;
In total 4 binding sites of Calcium where determined in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas, PDB code: 6dbl:
Jump to Calcium binding site number: 1; 2; 3; 4;
Calcium binding site 1 out of 4 in 6dbl
Go back to
Calcium binding site 1 out
of 4 in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
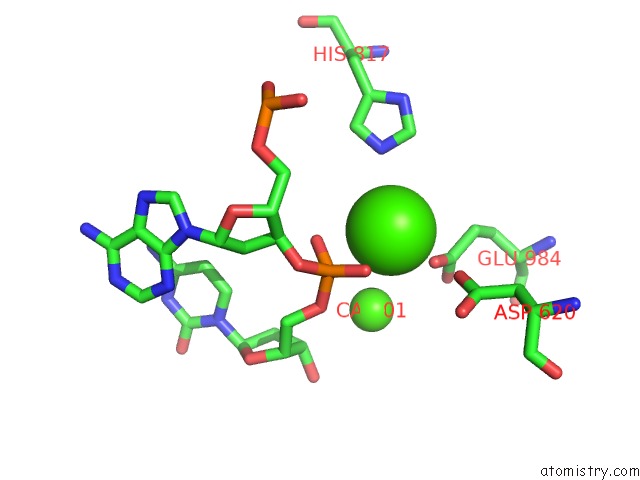
Mono view
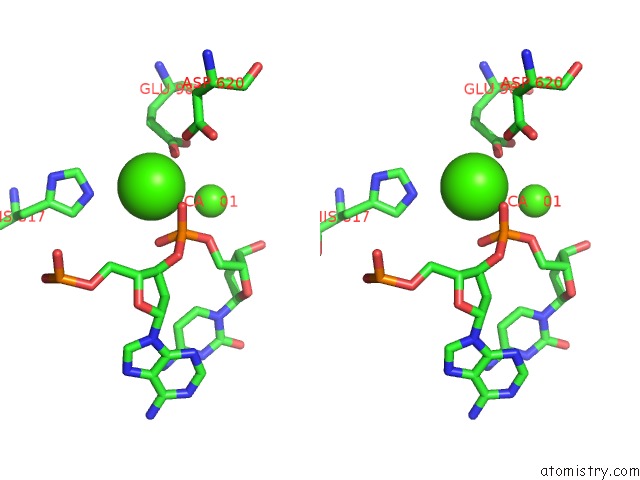
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas within 5.0Å range:
|
Calcium binding site 2 out of 4 in 6dbl
Go back to
Calcium binding site 2 out
of 4 in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
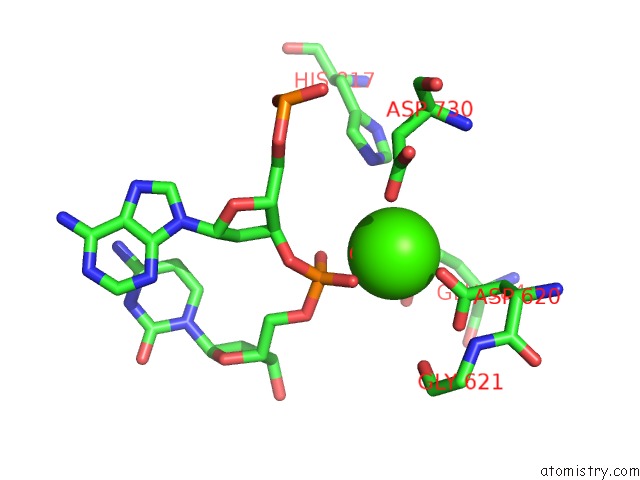
Mono view
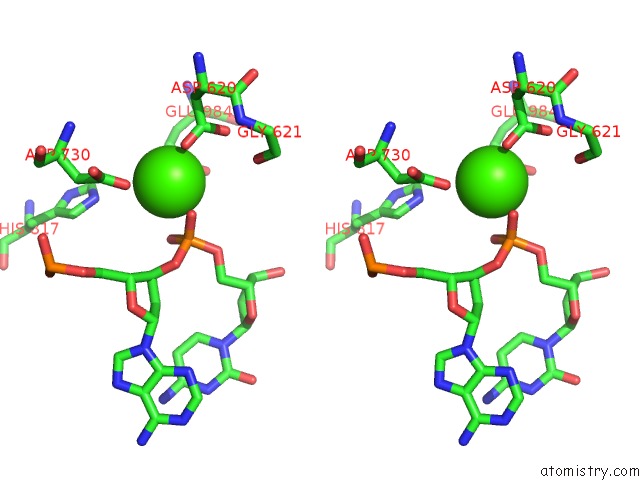
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 2 of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas within 5.0Å range:
|
Calcium binding site 3 out of 4 in 6dbl
Go back to
Calcium binding site 3 out
of 4 in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
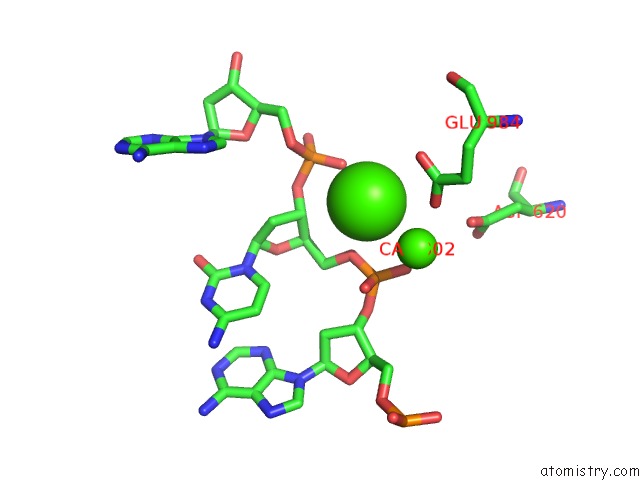
Mono view
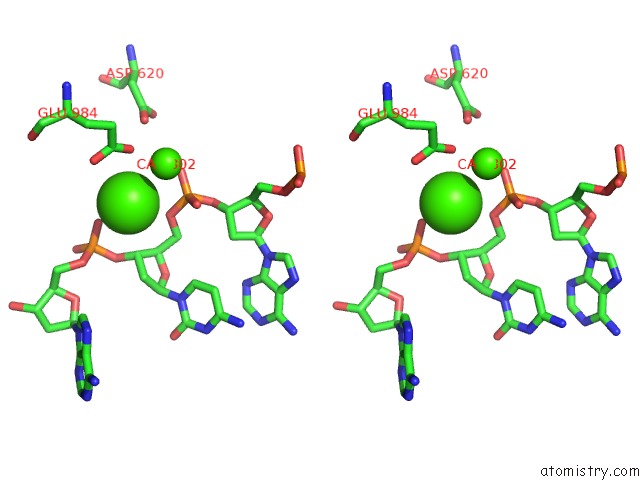
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 3 of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas within 5.0Å range:
|
Calcium binding site 4 out of 4 in 6dbl
Go back to
Calcium binding site 4 out
of 4 in the Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas
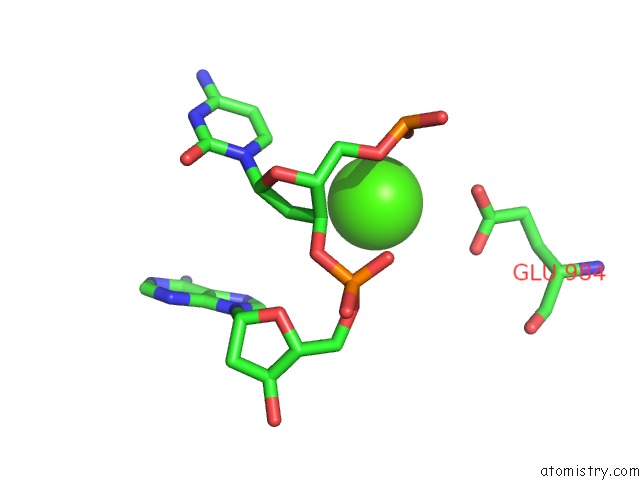
Mono view
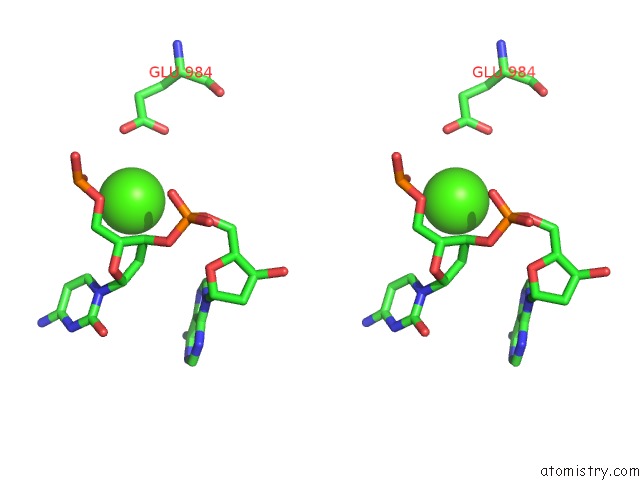
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 4 of Cryo-Em Structure of Rag in Complex with 12-Rss and 23-Rss Substrate Dnas within 5.0Å range:
|
Reference:
H.Ru,
W.Mi,
P.Zhang,
F.W.Alt,
D.G.Schatz,
M.Liao,
H.Wu.
Dna Melting Initiates the Rag Catalytic Pathway. Nat. Struct. Mol. Biol. V. 25 732 2018.
ISSN: ESSN 1545-9985
PubMed: 30061602
DOI: 10.1038/S41594-018-0098-5
Page generated: Mon Jul 15 17:48:56 2024
ISSN: ESSN 1545-9985
PubMed: 30061602
DOI: 10.1038/S41594-018-0098-5
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF