Calcium »
PDB 6hi0-6i1q »
6hr5 »
Calcium in PDB 6hr5: Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila
Protein crystallography data
The structure of Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila, PDB code: 6hr5
was solved by
T.Roret,
A.Prechoux,
M.Czjzek,
G.Michel,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 44.61 / 2.91 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 52.879, 48.871, 92.651, 90.00, 105.64, 90.00 |
| R / Rfree (%) | 32.2 / 36.1 |
Calcium Binding Sites:
The binding sites of Calcium atom in the Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila
(pdb code 6hr5). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total only one binding site of Calcium was determined in the Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila, PDB code: 6hr5:
In total only one binding site of Calcium was determined in the Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila, PDB code: 6hr5:
Calcium binding site 1 out of 1 in 6hr5
Go back to
Calcium binding site 1 out
of 1 in the Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila
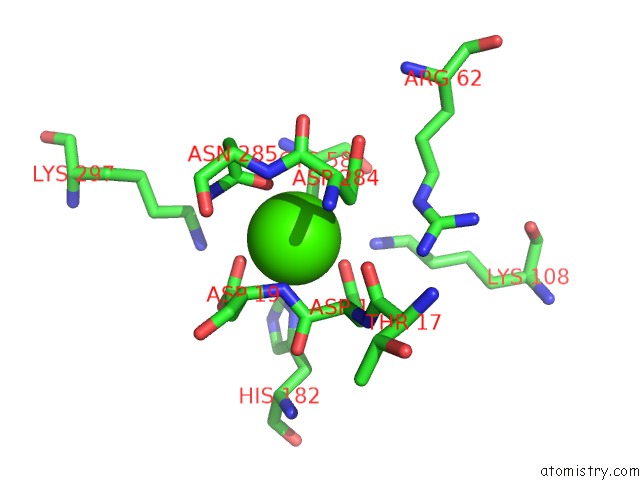
Mono view
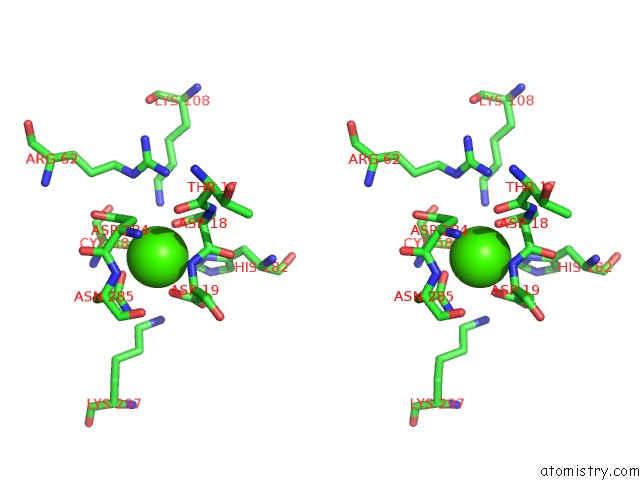
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Structure of the S1_25 Family Sulfatase Module of the Rhamnosidase FA22250 From Formosa Agariphila within 5.0Å range:
|
Reference:
L.Reisky,
A.Prechoux,
M.K.Zuhlke,
M.Baumgen,
C.S.Robb,
N.Gerlach,
T.Roret,
C.Stanetty,
R.Larocque,
G.Michel,
T.Song,
S.Markert,
F.Unfried,
M.D.Mihovilovic,
A.Trautwein-Schult,
D.Becher,
T.Schweder,
U.T.Bornscheuer,
J.H.Hehemann.
A Marine Bacterial Enzymatic Cascade Degrades the Algal Polysaccharide Ulvan. Nat.Chem.Biol. V. 15 803 2019.
ISSN: ESSN 1552-4469
PubMed: 31285597
DOI: 10.1038/S41589-019-0311-9
Page generated: Wed Jul 9 14:41:59 2025
ISSN: ESSN 1552-4469
PubMed: 31285597
DOI: 10.1038/S41589-019-0311-9
Last articles
Ca in 7M66Ca in 7M8C
Ca in 7M8B
Ca in 7M8T
Ca in 7M8A
Ca in 7M89
Ca in 7M88
Ca in 7M87
Ca in 7M86
Ca in 7M85