Calcium »
PDB 6kzo-6ljf »
6lgc »
Calcium in PDB 6lgc: Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin
Protein crystallography data
The structure of Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin, PDB code: 6lgc
was solved by
T.Miyazaki,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 48.97 / 1.90 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 65.634, 146.750, 154.025, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.3 / 19.7 |
Other elements in 6lgc:
The structure of Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin also contains other interesting chemical elements:
| Magnesium | (Mg) | 3 atoms |
Calcium Binding Sites:
The binding sites of Calcium atom in the Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin
(pdb code 6lgc). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total 2 binding sites of Calcium where determined in the Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin, PDB code: 6lgc:
Jump to Calcium binding site number: 1; 2;
In total 2 binding sites of Calcium where determined in the Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin, PDB code: 6lgc:
Jump to Calcium binding site number: 1; 2;
Calcium binding site 1 out of 2 in 6lgc
Go back to
Calcium binding site 1 out
of 2 in the Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin
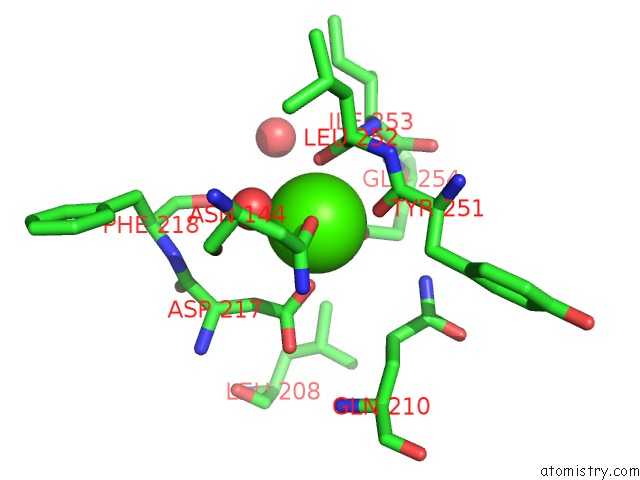
Mono view
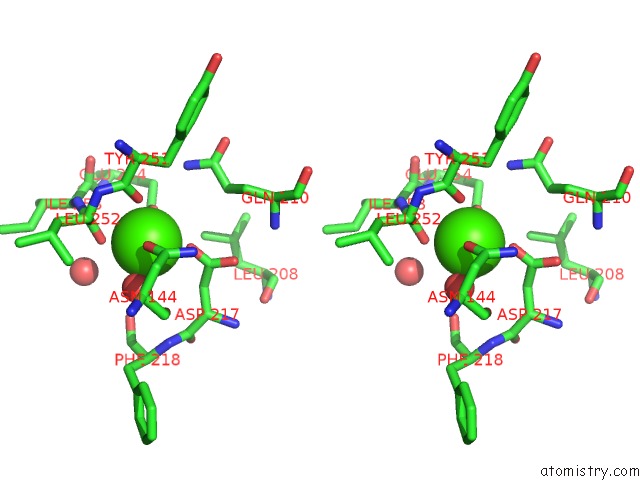
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin within 5.0Å range:
|
Calcium binding site 2 out of 2 in 6lgc
Go back to
Calcium binding site 2 out
of 2 in the Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin
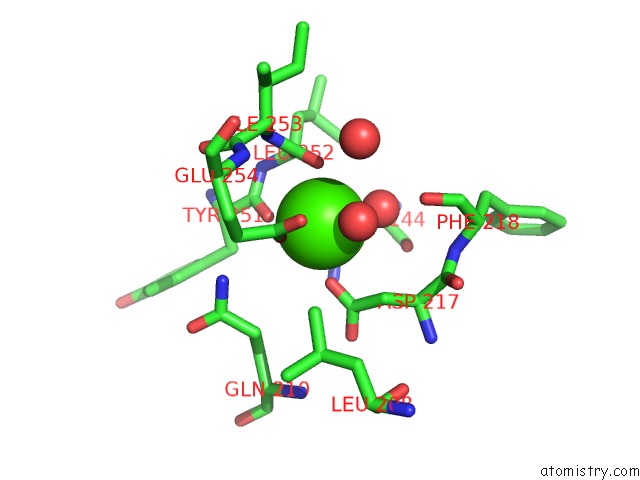
Mono view
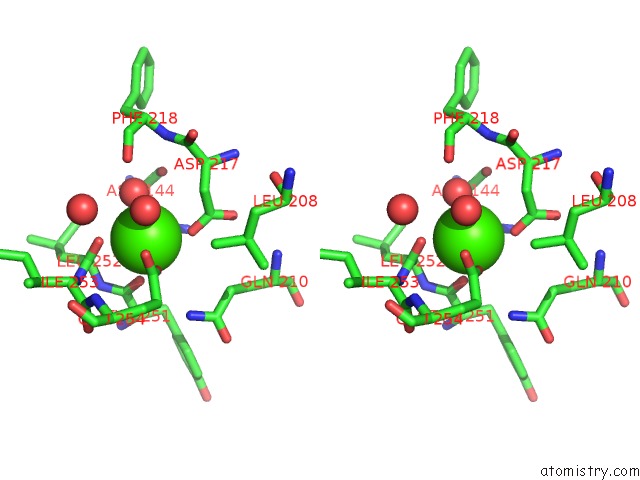
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 2 of Bombyx Mori GH13 Sucrose Hydrolase Complexed with 1-Deoxynojirimycin within 5.0Å range:
|
Reference:
T.Miyazaki,
E.Y.Park.
Structure-Function Analysis of Silkworm Sucrose Hydrolase Uncovers the Mechanism of Substrate Specificity in GH13 Subfamily 17EXO-Alpha-Glucosidases. J.Biol.Chem. 2020.
ISSN: ESSN 1083-351X
PubMed: 32381508
DOI: 10.1074/JBC.RA120.013595
Page generated: Wed Jul 9 15:51:50 2025
ISSN: ESSN 1083-351X
PubMed: 32381508
DOI: 10.1074/JBC.RA120.013595
Last articles
Cl in 5L79Cl in 5L77
Cl in 5L6J
Cl in 5L74
Cl in 5L6Q
Cl in 5L6C
Cl in 5L69
Cl in 5L6B
Cl in 5L6A
Cl in 5L66