Calcium »
PDB 6ai0-6b40 »
6avz »
Calcium in PDB 6avz: Crystal Structure of the Hopq-CEACAM3 Wt Complex
Protein crystallography data
The structure of Crystal Structure of the Hopq-CEACAM3 Wt Complex, PDB code: 6avz
was solved by
D.A.Bonsor,
E.J.Sundberg,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 103.26 / 2.05 |
| Space group | P 2 2 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 39.479, 103.263, 112.565, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 20.3 / 25 |
Calcium Binding Sites:
The binding sites of Calcium atom in the Crystal Structure of the Hopq-CEACAM3 Wt Complex
(pdb code 6avz). This binding sites where shown within
5.0 Angstroms radius around Calcium atom.
In total 2 binding sites of Calcium where determined in the Crystal Structure of the Hopq-CEACAM3 Wt Complex, PDB code: 6avz:
Jump to Calcium binding site number: 1; 2;
In total 2 binding sites of Calcium where determined in the Crystal Structure of the Hopq-CEACAM3 Wt Complex, PDB code: 6avz:
Jump to Calcium binding site number: 1; 2;
Calcium binding site 1 out of 2 in 6avz
Go back to
Calcium binding site 1 out
of 2 in the Crystal Structure of the Hopq-CEACAM3 Wt Complex
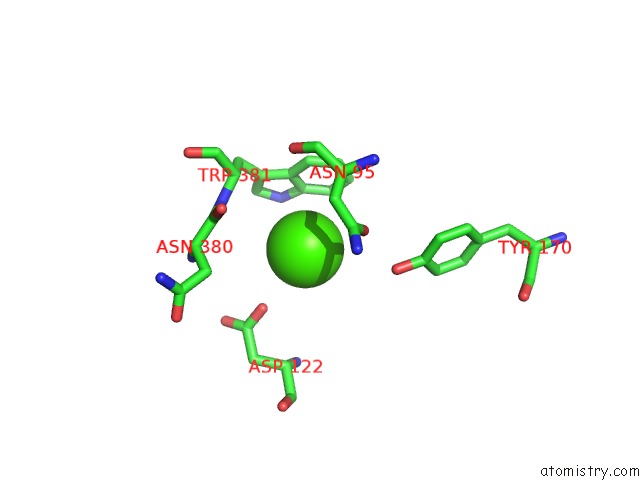
Mono view
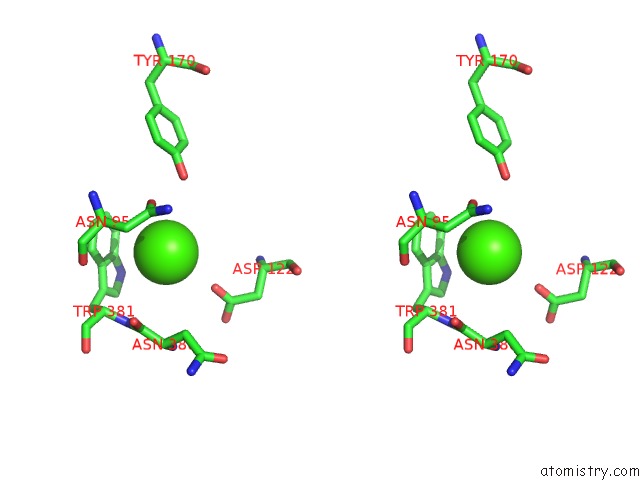
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 1 of Crystal Structure of the Hopq-CEACAM3 Wt Complex within 5.0Å range:
|
Calcium binding site 2 out of 2 in 6avz
Go back to
Calcium binding site 2 out
of 2 in the Crystal Structure of the Hopq-CEACAM3 Wt Complex
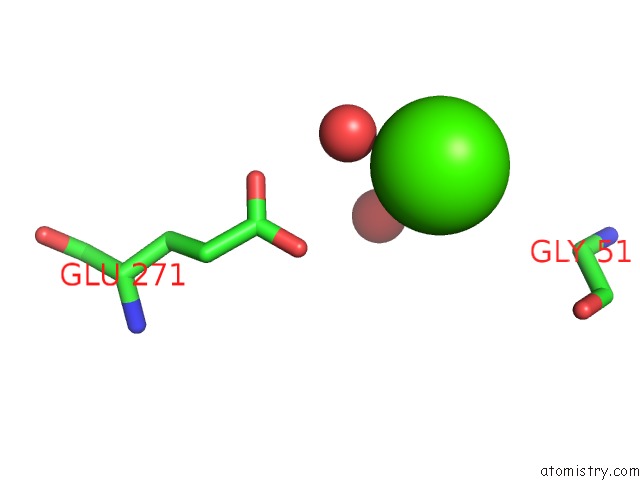
Mono view
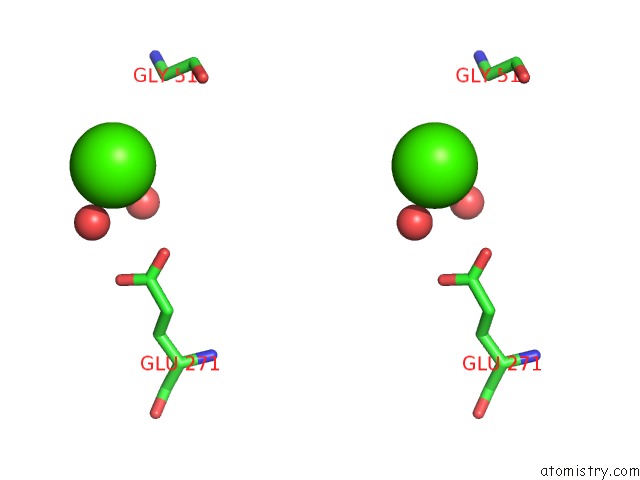
Stereo pair view

Mono view

Stereo pair view
A full contact list of Calcium with other atoms in the Ca binding
site number 2 of Crystal Structure of the Hopq-CEACAM3 Wt Complex within 5.0Å range:
|
Reference:
D.A.Bonsor,
Q.Zhao,
B.Schmidinger,
E.Weiss,
J.Wang,
D.Deredge,
R.Beadenkopf,
B.Dow,
W.Fischer,
D.Beckett,
P.L.Wintrode,
R.Haas,
E.J.Sundberg.
Thehelicobacter Pyloriadhesin Protein Hopq Exploits the Dimer Interface of Human Ceacams to Facilitate Translocation of the Oncoprotein Caga. Embo J. V. 37 2018.
ISSN: ESSN 1460-2075
PubMed: 29724755
DOI: 10.15252/EMBJ.201798664
Page generated: Wed Jul 9 12:34:17 2025
ISSN: ESSN 1460-2075
PubMed: 29724755
DOI: 10.15252/EMBJ.201798664
Last articles
Fe in 2YXOFe in 2YRS
Fe in 2YXC
Fe in 2YNM
Fe in 2YVJ
Fe in 2YP1
Fe in 2YU2
Fe in 2YU1
Fe in 2YQB
Fe in 2YOO